"Can genes determine your destiny? When it comes to genetics, we are surrounded by ignorance, fake news, myths, and disinformation. It’s time you knew the truth! In this interview, Gene Heritage Founder and managing director Castedo Ellerman discusses some misconceptions about genetics and explains why you should never expect deterministic answers from DNA analysis." ... Read the full interview on DNAweekly.
Blog Posts
- 2020-06-25 Breaking Old Myths About Genetics And DNA
- 2019-03-09 How real are DNA test results?
- 2018-11-21 Food and Your Genes
- 2018-09-03 What Percentage of DNA Did I Inherit From My Grandparents?
- 2018-09-02 What Percentage of DNA Did I Inherit From My Parents?
- 2018-08-28 How Does ABCC11 Influence My Earwax Type?
- 2018-08-23 Does ABCC11 Influence My Body Odor?
- 2018-05-23 What is Raw DNA Data?
- 2018-05-16 How Do I Use Raw DNA Data?
- 2018-05-11 What Else Can I Do with My Raw DNA Data?
How real are DNA test results?
Many services today use personal DNA to generate a wide variety of reports. But how real are reported results? And why do so many people see results that seem wrong? Recognizing that reported results are predictions helps answer these questions. Report services look at personal DNA and then predict something. All attempts to predict carry two risks: honest misprediction and pseudo prediction. We can judge how real report results are by the degree of both these risks.
Honest Misprediction
Most genetic reports use real science and are not completely made up. That scientific prediction makes some mispredictions should be no surprise. But these honest mispredictions can occur surprisingly often in genetic reports. Individual reported results need not be rarely incorrect to be scientific.
Take, for example, a car inspection service. It provides a personalized report based on your car instead of your DNA. A car inspection might report ‘at risk of brake failure’. There is a real scientific basis for this prediction based on the physical condition of the brakes. But the warning will still prove to be a misprediction for many unwise drivers who ignore the warning (the lucky ones).
Pseudo Prediction
Some reported results are too ambiguous and too amenable to wide interpretation. These predictions apply to almost all individuals no matter what their DNA. The results might be true, but are they really reporting something about your DNA if they apply to everyone’s DNA?
Imagine another car inspection service that reports your car ‘will get people moving’. The assertion "will get people moving" is easily interpreted as correct with ANY car. The inspection is a phony act of prediction. The report result is true, but it’s a pseudo prediction. Services can reduce the degree of honest misprediction by resorting to pseudo prediction. Reports will still feel accurate thanks to the Barnum effect (also known as the Forer effect). In another post to follow, you can see more examples of pseudo predictions and the Barnum effect.

Predictability Score
So, how real are reported results? That depends on the combined degree of both honest misprediction and pseudo prediction. Gene Heritage reports include a predictability score to help readers appreciate this combined degree. Three simple grades of ‘Highly Predictable’, ‘Fairly Predictable’, or ‘Barely Predictable’ summarize this score. Results at risk of being pseudo predictions only appear with a "Barely Predictable" score. A "Highly Predictable" score is only achieved with a small amount of honest misprediction. In a post to follow, see predictability scores for some familiar example traits. The next post will dive into the theory and mathematical details behind predictability scores.
Food and Your Genes

What better way to spend this year’s Thanksgiving dinner than to get into an epic family fight over the current state of politics! Well…maybe not. If you’re looking for something to talk about that won’t ruffle any feathers (other than the turkey’s), how about the topic of food and your genes? Here are some examples of how genes influence your perception of different food and beverages:
TAS2R38 Gene — influences your sensitivity to bitterness in certain naturally occurring chemicals in red wine and cruciferous vegetables, such as Brussels sprouts and broccoli.
OR6B2 Gene — influences smell sensitivity to a chemical found in many foods and beverages including beer, Gouda cheese, Thai fish sauce, dark chocolate, tea infusions, and truffles.
TAS2R31 Gene Region — influences bitter taste sensitivity to saccharin, an artificial sweetener used in drinks, candies, and medicines. It’s a primary ingredient in Sweet ‘N Low. The TAS2R31 region also influences sensitivity to caffeine and quinine, as well as a chemical found in artichokes.
TAS1R3 Gene Region — influences taste sensitivity to sugars, like sucrose, glucose, and fructose. For humans and other primates, the ability to taste sweetness is an important trait as our diet relies largely upon fruit and other plant foods containing sugar. By contrast, cats and many carnivorous animals have lost their sweet receptor over the course of evolution.
TAS1R1 Gene — influences taste sensitivity to umami, the mysterious "fifth" basic taste, along with sweetness, sourness, bitterness, and saltiness. Umami means “pleasant savory taste” in Japanese. It’s typically described as “savory,” “meaty,” and “brothy,” as well as having a mouthwatering and coating sensation over the tongue. It’s found in glutamate, a naturally-occurring compound in meat broths, fermented products, tomatoes, cheese, and many other foods.
It’s important not to overstate the influence of these genes; most only have a moderate to minor influence on your taste and smell sensitivity. Still, it’s interesting to see how genes were passed down in your family and to learn about their ancient origins. Get a Gene Heritage report to compare your genes with other family members and to build an inheritance tree. You may or may not agree with your Uncle Bob’s views on politics, but it may be consoling—and healing—to find you share the same TAS1R1 gene!
What Percentage of DNA Did I Inherit From My Grandparents?

As discussed in the previous blog, parents pass down a random half of their DNA to their children. Whether that random half has more DNA from a grandfather or grandmother is also random. So even though children get 49-51% of their DNA from each parent, the amount of DNA they inherit from each grandparent can be markedly different than 25%. If your grandparents get testy about this, remind them it’s not a competition!
Gene Heritage’s Grandchild Report calculates what percentage of DNA an individual inherited from each grandparent. In this example Grandchild Report, Jean Mendel inherited 30% of his DNA from his paternal grandmother Greta and only 19% from his paternal grandfather Gregorious. As you can see from the report’s donut chart, even though we inherit on average about 25% of our DNA from each grandparent, the actual percentages can be strikingly different.
What Percentage of DNA Did I Inherit From My Parents?
How much DNA did you inherit from your mother and father? If you guessed 50% from each parent... well, you’re only half right. While women do inherit 50% of their DNA from each parent, men inherit about 51% from their mother and only 49% from their father.
For all you men out there, is this proof you really are a mama’s boy? Why the discrepancy?
To answer this question, first a little 101 in genetics: all humans, both male and female, inherit 23 chromosome pairs from their parents, for a total of 46 chromosomes. Half of each pairing comes from an individual’s mother and half from the father. In 22 of these chromosome pairings (known as “autosomes”), each half of the pairing is roughly the same length and size.

A 23rd chromosome pairing consists of two “allosomes,” more commonly known as “sex chromosomes.” Sex chromosomes can either be an X or Y. Unlike the 22 autosomes, X and Y chromosomes are NOT the same size.
The sex chromosome pairing for a man is XY, meaning all you fellas out there inherited one X from your mother and one Y from your father. Women, on the other hand, have a sex chromosome pairing of XX, inheriting one X chromosome from each parent.
The Y chromosome found in males is about one-third the size of an X chromosome—and contains significantly less DNA. An X chromosome has HUNDREDS more genes than a Y chromosome. It’s this discrepancy in the size of X and Y chromosomes that accounts for why men inherit 51% of their DNA from their mothers and only 49% from their fathers.
With a Gene Heritage Parent-Child Report, you can see which genes have been passed down from a parent to a child.
As a side note, mitochondrial DNA is passed down exclusively along the maternal line (from mothers to children), but only accounts for a piddling 0.0003% of your DNA . Still, it’s just a bit more proof that women have a leg up on all you men out there. But does size really matter?
How Does ABCC11 Influence My Earwax Type?
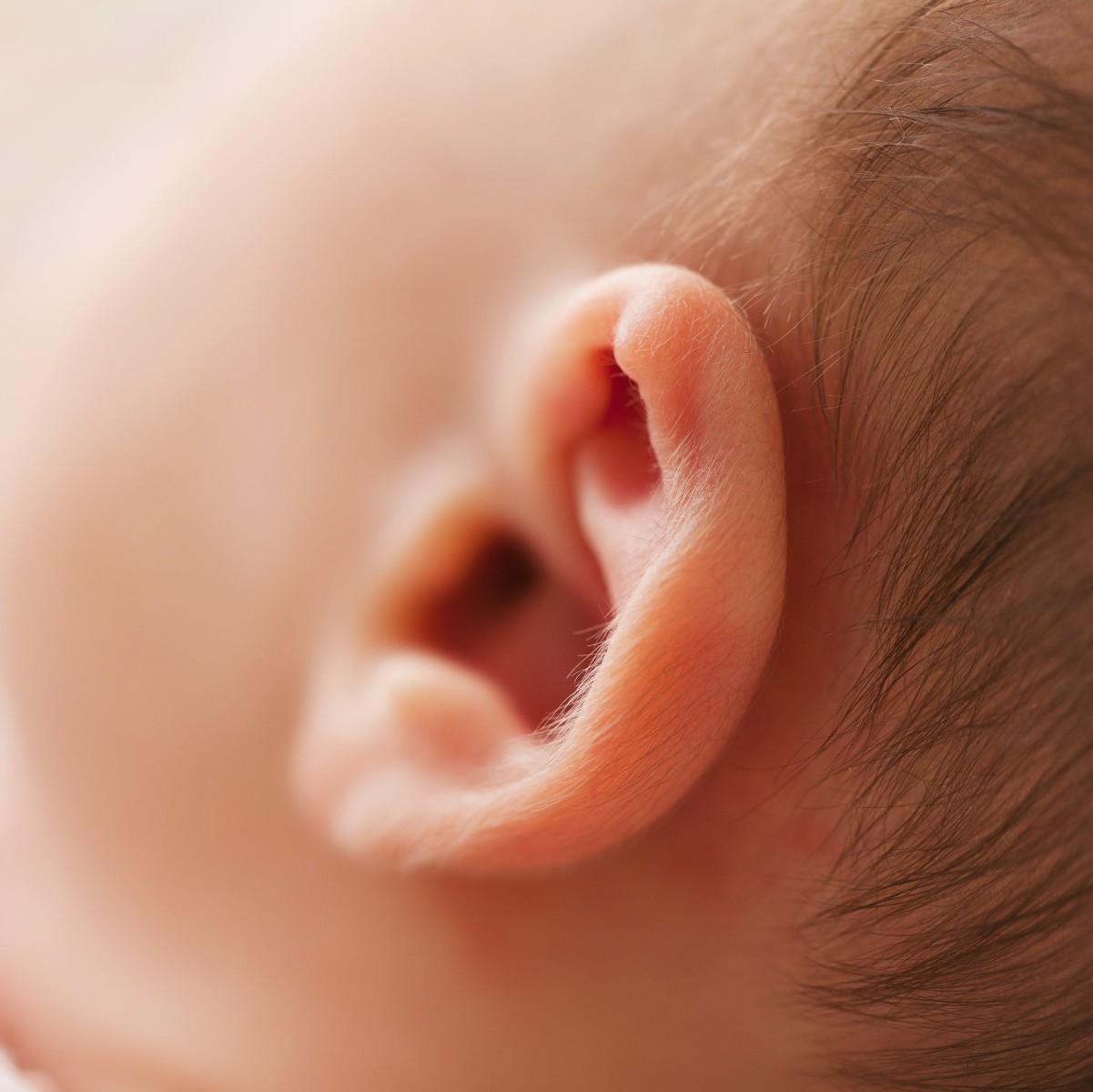
In our last blog entry, we discussed ABCC11’s relatively minor influence on body odor. But did you know ABCC11 has a much stronger influence on whether you have dry or wet earwax? If you aren’t grossed out yet, read on …
As with armpit odor, ABCC11 influences specialized skin glands, but in this case, the glands control earwax type. Dry, flaky earwax is more prevalent in Asian populations, whereas moist, brownish earwax is more common in people of African and European ancestry.
According to one study, the mutation for the “Dry” allele first appeared around 40,000 years ago in Northeast Asia, spreading to other parts of the continent. It’s theorized natural selection favored the mutation in the cold climes of Eastern Asia.
Don’t knock earwax: as gross as it might seem, earwax (or “cerumen”) aids in the cleaning and lubrication of your ear canals. It also provides protection against bacteria, fungi, insects, and water.
Does ABCC11 Influence My Body Odor?
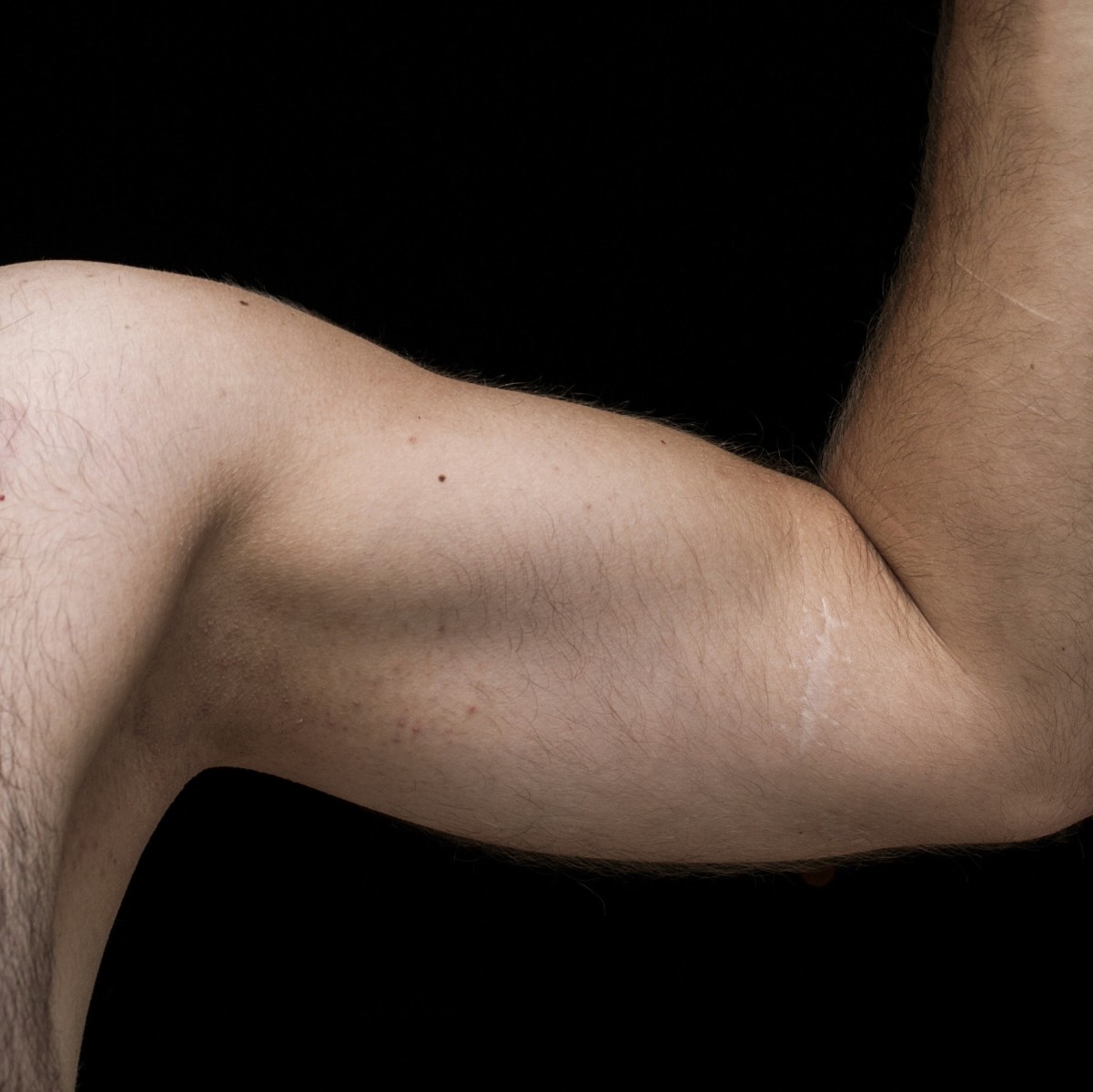
Let’s face it: from time to time, we all sniff our very own armpits to take a whiff of our axillary odor. Sometimes our underarms pass the smell test; other times we recoil in shock. Is there a genetic component to armpit odor? The answer is yes. But before you run out to Costco to buy deodorant in bulk, put yourself at ease by reading the rest of this blog.
Located on chromosome 16 is a gene called ABCC11 that influences armpit odor. People with the “Wet” allele of ABCC11 tend to produce more body odor than those with only “Dry” alleles. The “Dry” allele is much more prevalent in Asia — it’s most common in Korea, China, Mongolia, and western Japan.
The “Wet” allele, on the other hand, is most common in African and European populations. While people with African or European ancestry may be predisposed to more body odor than East Asians, ABCC11’s influence on armpit odor is only minor. The ABCC11 gene actually has a much more direct and major influence on your earwax type.
Many non-genetic factors influence armpit odor in addition to your genes, especially personal hygiene. Gender, disease, stress, and diet may also influence your axillary odor. So before you go on your next date or job interview, please don’t overthink ABCC11’s influence. You probably smell just fine.
What is Raw DNA Data?
When you mail in your saliva sample to AncestryDNA™ or another DNA collector they use a nifty device called a microarray to map your genome. They don’t look at your entire genetic code, because our genomes are by and large 99.9% the same across all humans (I bet you feel soooo special now). Instead, they focus on the 0.1% of known common genetic variation between us. These variations, also known as markers, or SNPs (“snips”), account for most of the human genetic variations you observe. They account for differences in eye color, skin pigmentation, and many other traits.
“Raw DNA data” sounds like a mouthful but it’s simply a computer file that lists out these SNPs, along with other useful information which we’ll get to later. It’s a text file you can save to your desktop, or double-click to open.
AncestryDNA™ and 23andMe provide raw data in a .txt format (compressed during download as a .zip); FTDNA provides it as a .CSV file (compressed as a .gz).
More about Your Raw DNA Data
While raw DNA data doesn’t contain your entire genome, it does include, oh, just several hundred thousand SNPs. AncestryDNA™’s and FTDNA’s raw data files include about 700,000 markers; 23andMe’s include anywhere from about 600,000 to 1 million markers depending on when you were tested. Here’s an approximate SNP count for the various DNA collectors:
| Approx. # SNPs | DNA Collector |
|---|---|
| 701,480 | AncestryDNA v1 (pre-May 2016) |
| 668,940 | AncestryDNA v2 |
| 571,430 | 23andMe v2 (really old) |
| 949,460 | 23andMe v3 (pre-Nov 2013) |
| 552,540 | 23andMe v4 (pre-Sep 2017) |
| 620,290 | 23andMe v5 |
| 700,000 | FamilyTreeDNA (various versions) |
| 720,710 | MyHeritage v1 & v2 |
| 606,130 | Living DNA v1 |
| 536,070 | Genes for Good v1 |
| 178,600 | Geno 2.0 |
If I had a dollar for every SNP I have…
DNA collectors only use a very small portion of your DNA to generate their results. The big draw of third-party tools like Gene Heritage is they give you a second chance to mine your unused raw DNA data for even more information about yourself. The difference in SNP counts between DNA collectors explains why coverage may vary with your third-party tool.
Typically all you need to do with your raw DNA data is download it from your DNA collector and upload it to a third-party party tool. But if you want to get in touch with your inner geek by ogling a long list of your single nucleotide polymorphisms, or SNPs, open your raw DNA file in a text editor and you’ll see something like this:
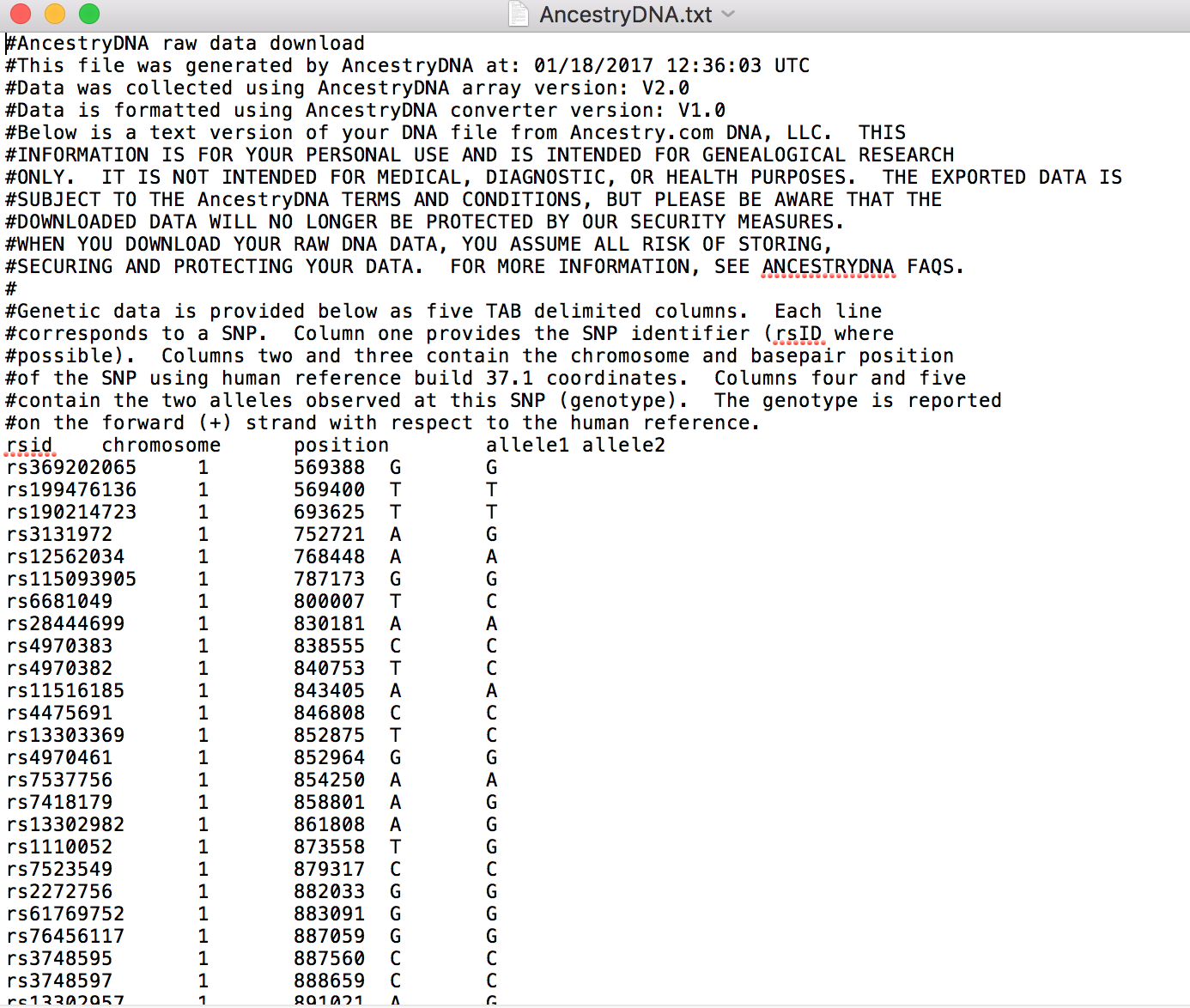
This is a screenshot from my very own AncestryDNA™ raw DNA data (please don’t use it to clone me; I’m not that special, trust me). Raw DNA from other collectors looks very similar, typically with five columns corresponding to:
- Your SNPs (each identified by a code called a rsID number)
- The chromosomes on which each SNP is located
- The position of each SNP on the chromosome
- Your two alleles for each SNP (one comes from your father, one from your mother). Possible allele letters are A (adenine), C (cytosine), G (guanine), T (thymine), or 0 (for missing data).
In this screenshot, the highlighted line corresponds to a variation influencing my eye color.
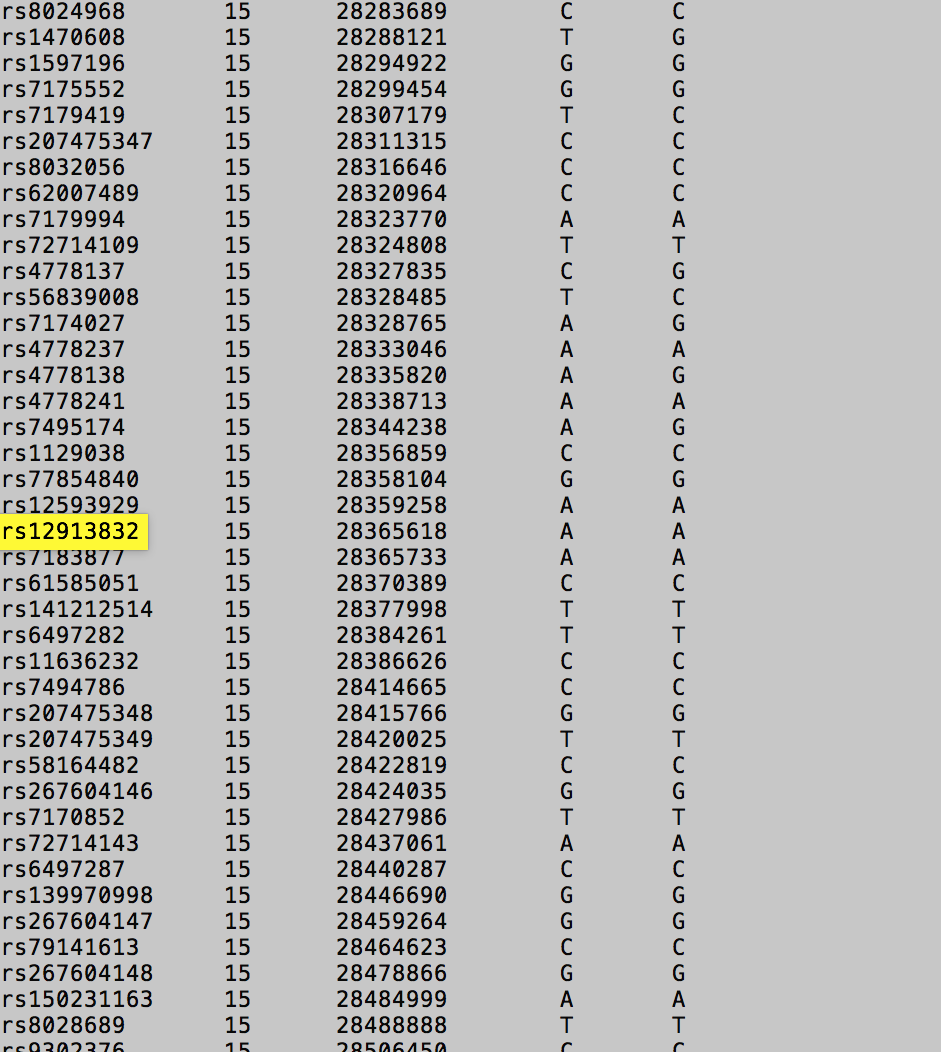
From this highlighted line I can see that:
- The rsID of the SNP is rs12913832
- The SNP is located on my 15th chromosome (each human has 23 chromosomes)
- I inherited one A allele from my mother and another A allele from my father.
How Do I Use Raw DNA Data?
How Do I Use Third-Party Raw DNA Tools?
To use a third-party tool, first download your raw DNA data from your DNA collector, then simply upload it to the third-party tool. Some tools only accept raw DNA from one or two DNA collectors; others like Gene Heritage are compatible with raw DNA from multiple sources. Click here to see which tools accept raw DNA transfers from your DNA collector.
How Do I Get My Raw DNA Data?
If you’ve sent in a saliva sample to your DNA collector and received your results, you already have access to your very own raw DNA file. Login to the website of your DNA collector to download it for free. Here are instructions on how to download your raw DNA.
Please read your DNA collector’s Terms of Service and Privacy Statement before downloading; most don’t want you to get mad at them if third-party tools reveal scary information that keeps you up all night.
In all seriousness, some third-party tools make dubious or uninformed claims that could be misleading or cause anxiety. It’s important to keep in mind that there are a whole bunch of other influences on your traits aside from genes, including dietary, microbial, and lifestyle factors. Talk to a health care provider or genetic counselor if you seek professional health care assistance.
What Else Can I Do with My Raw DNA Data?
It wasn’t pretty, but you managed to spit a big gob of saliva into a vial and mail it off to AncestryDNA™, 23andMe, FTDNA or some other DNA collector. After waiting on pins and needles for a month or two, you finally got your results.
Now what? What else can you do with your DNA results?
With AncestryDNA™, 23andMe, FTDNA, and other DNA collectors you get a file known as "raw DNA data" that third-party tools like Gene Heritage can use to tell you even more about yourself.
Why Use Third-Party Raw DNA Tools?
Third-party raw DNA tools use your raw data to:
- Tell you more about your genes and traits
- Dig deeper into your genetic genealogy
- Match you with long-lost relatives
- Tell you more about your health
Some third-party tools charge a small fee (like Gene Heritage or Promethease); others are pretty expensive. Click here for a list of raw DNA tools.